
(A) Phylogenetic tree of Egyptian rousette IFN-ω proteins. We characterized the antiviral potency and efficacy of these recombinant proteins against VSV and against MARV, and examined the downstream ISGs they induce.Ĭomparison of five clades of Egyptian rousette IFN-ω proteins. To begin to understand the role of these genes in the immune response to viruses in bats, we synthesized and purified recombinant Egyptian rousette IFN-ω proteins. The ISGs induced specifically by these IFN-ω proteins, however, are not known. Multiple subtypes of porcine IFN-ω are expressed after viral infection and have dramatic differences in activity despite very few single nucleotide polymorphisms ( 23).

In humans and other species, IFN-ω is induced by viral infection and has potent antiviral activity against various RNA viruses, including vesicular stomatitis virus (VSV), bovine viral diarrhea virus, yellow fever virus, West Nile virus, and influenza A virus ( 20– 24). The functional relevance of the expansion is not known. Whereas, humans have a single IFNW gene, almost half of the Egyptian rousette IFN genes belong to the IFN-ω subtype. aegyptiacus), an asymptomatic host of Marburg virus (MARV) ( 19). We have previously shown that the type I IFN locus is expanded in the Egyptian rousette ( R.
More recently, bat IFN responses have been explored using recombinant bat IFN-α or –β, but additional bat IFN subtypes remain poorly characterized ( 16– 18). Due to the lack of bat-derived IFNs, much of the work to analyze IFN responses in bats thus far has been done using universal interferon (UIFN a pan-species type I IFN derived from two human IFN-α subtypes) or cell culture medium from stimulated bat cells as a surrogate for authentic bat IFN ( 13– 15). The importance of type I IFNs in innate antiviral responses and in bridging innate and adaptive immune responses has sparked interest in exploring this pathway in several bat species. As a result, differences in pathogen-specific antiviral effect are possible, depending on the amount and profile of ISGs induced by a particular IFN. Although the exact functional contribution for each IFN is not completely understood, differences in the interaction of various IFN subtypes with IFNAR1/2 are known to differentially induce downstream ISGs ( 10– 12). All type I IFNs bind to and signal through the same heterodimeric receptor complex IFNAR1/2, but both evolutionary analyses and functional studies suggest that multiple IFN subtypes make non-redundant contributions to immunity ( 6– 10). Type I IFNs are induced by the recognition of viral pathogen-associated molecular patterns (PAMPs), and act by inducing interferon stimulated genes (ISGs) that collectively contribute to an antiviral response ( 4, 5). Type I interferons (IFNs) are an important component of the early antiviral immune response, and make up a diversified multi-gene family, including subtypes like α, β, δ, ω, ε, and others ( 3). The adaptations (in host or virus) that allow bats to host emerging viruses without developing symptoms of disease are not yet known.

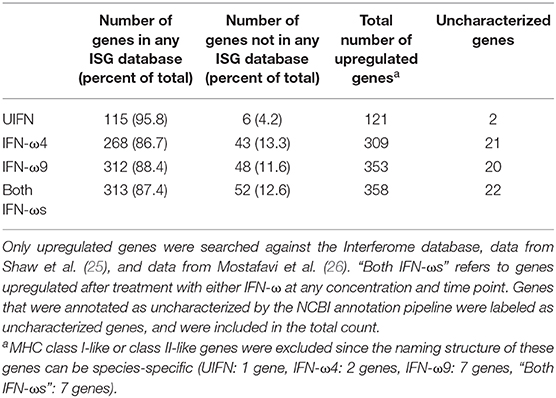
While humans develop severe and life-threatening illnesses from many of these viruses (e.g., henipaviruses, SARS and MERS coronaviruses, and filoviruses), bats show no symptoms of disease in natural or experimental infections ( 1, 2). About 13% of genes induced by IFN-ω treatment are not found in the Interferome or other ISG databases, indicating that they may be uniquely IFN-responsive in this bat.īats comprise about 20% of all classified mammal species with over 1,200 species and host a number of viruses known to cause severe disease in humans. Although the sets of genes induced by the two IFNs were largely overlapping, IFN-ω9 induced a more rapid and intense response than did IFN-ω4. Using RNA-Seq, we examined the transcriptional response induced by each protein. Both IFN-ω4 and IFN-ω9 showed antiviral activity against RNA viruses, including MARV, with IFN-ω9 being more efficient than IFN-ω4. To probe the biological implications of this expansion, we synthesized IFN-ω4 and IFN-ω9 proteins and tested their antiviral activity in Egyptian rousette cells. A striking difference to the human genome is that Egyptian rousettes have an expanded repertoire of IFNW genes. In contrast to humans, these bats do not become sick when infected with MARV. Egyptian rousette bats ( Rousettus aegyptiacus) are the natural reservoir hosts of Marburg virus (MARV). While the mechanisms underlying this ability to avoid sickness are not known, deep sequencing studies of bat genomes have uncovered genetic adaptations that may have functional importance in the antiviral response of these animals. Bats host a number of viruses that cause severe disease in humans without experiencing overt symptoms of disease themselves.
